The present investigation was carried out at the research farm of The Department of Genetics and Plant Breeding, Chaudhary Charan Singh Haryana Agricultural University, Hisar under irrigated condition and Regional Research Station, CCS Haryana Agricultural University, Bawal (Rewari) under rainfed conditions. The experiment was conducted during the kharif season, 2011. The seed material for the present study consisted of 24 genotypes (7 hybrids, 6 CMS lines, 6 B lines, 5 restorers) of pearl millet as given in Table 1. A set of 24 genotypes was evaluated in randomized block design (RBD) with three replications under irrigated conditions. The plot size was one row of 4 m length with row to row distance of 50 cm and plant to plant 15 cm. Recommended agronomic practices were followed to raise the crop. The observations were recorded on 5 plants in each replication at specified stages of crop growth period when the characters under study had full expression. Pooled data from both locations were used to classify the genotypes. Flow charts were also prepared depicting key diagnostic characters related to genotypes.
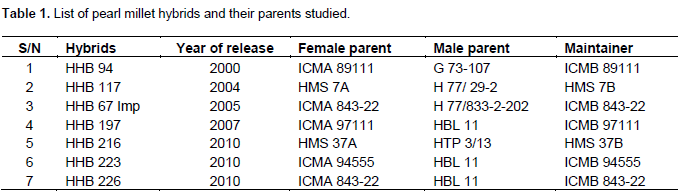
The identification of a pearl millet genotype is established by using a set of morphological characteristics. These characteristics are also useful to establish distinctness, uniformity and stability of the new variety, based on which the variety is given protection. In the present study, 28 morphological characteristics listed in the DUS test guidelines and morphological descriptors for pearl millet were explored for varietal description of pearl millet. Among the 24 pearl millet genotypes, considerable variation was observed between the genotypes for all the important attributes under study at both locations except anthocynin coloration of first leaf sheath, number of nodal tillers, spike girth, stem thickness, leaf mid rib colour and ligule hair. All the qualitative traits, as expected, were found similar over the locations (Kumar et al., 1993). The characterization of pearl millet hybrids and their parents under study is presented in Table 2. Key diagnostic characters of each genotype are assigned through flow chart in Figure 1.
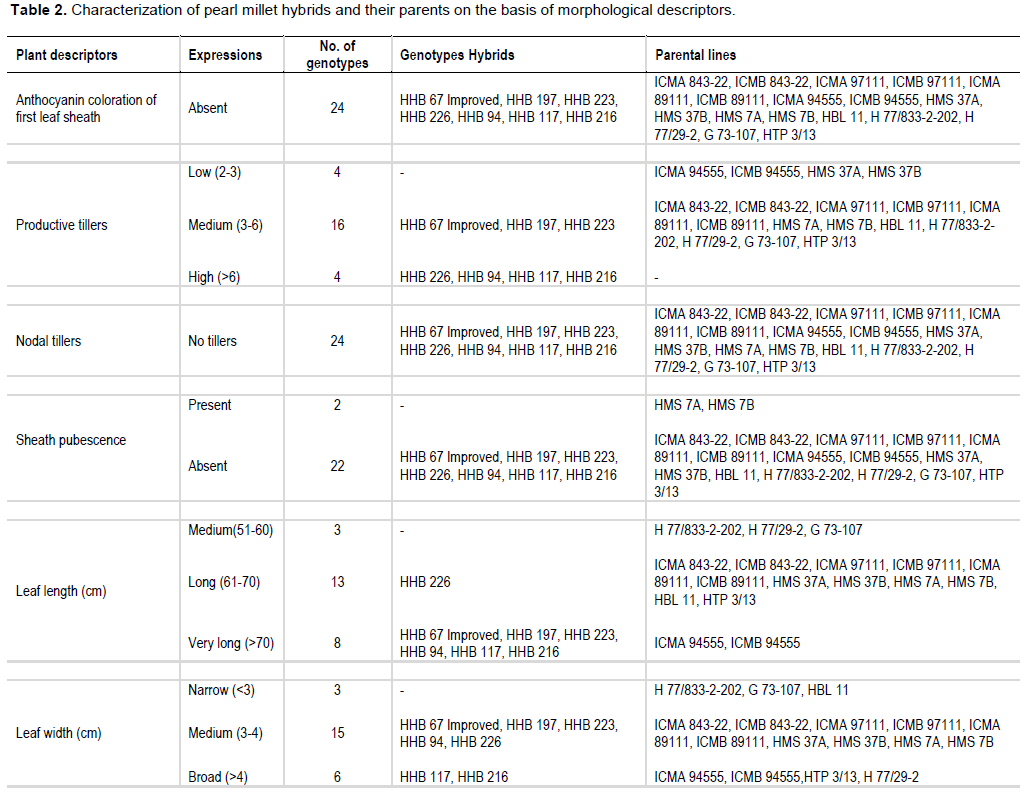
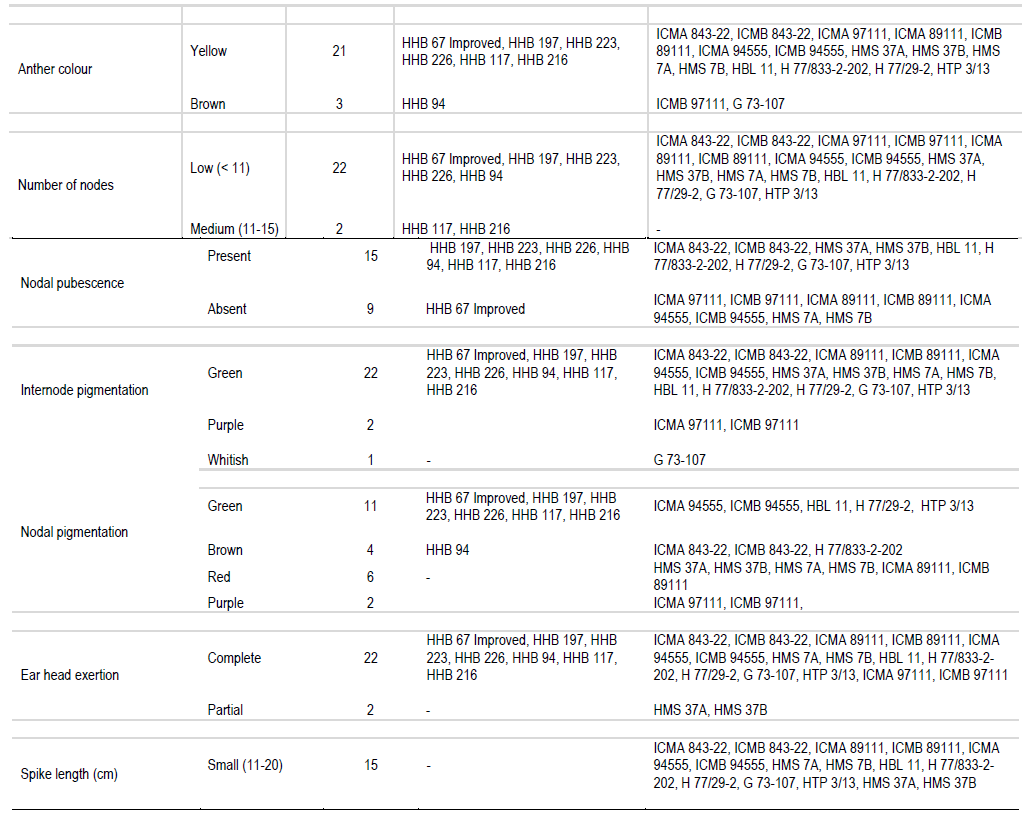
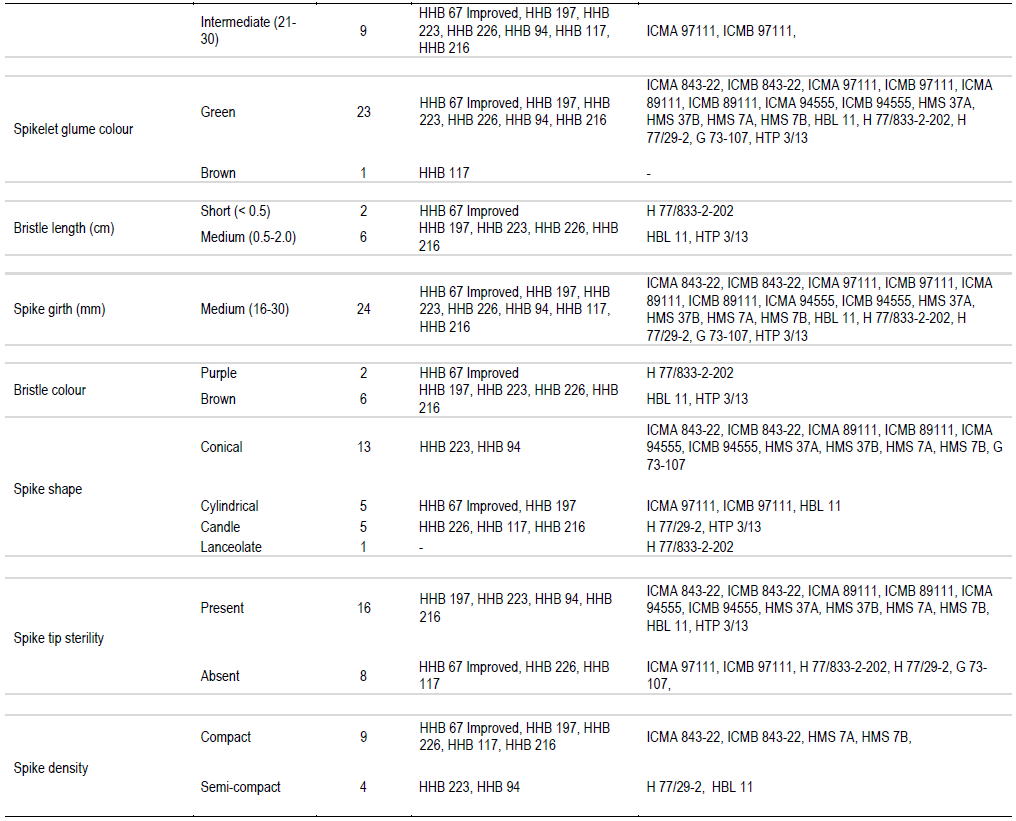
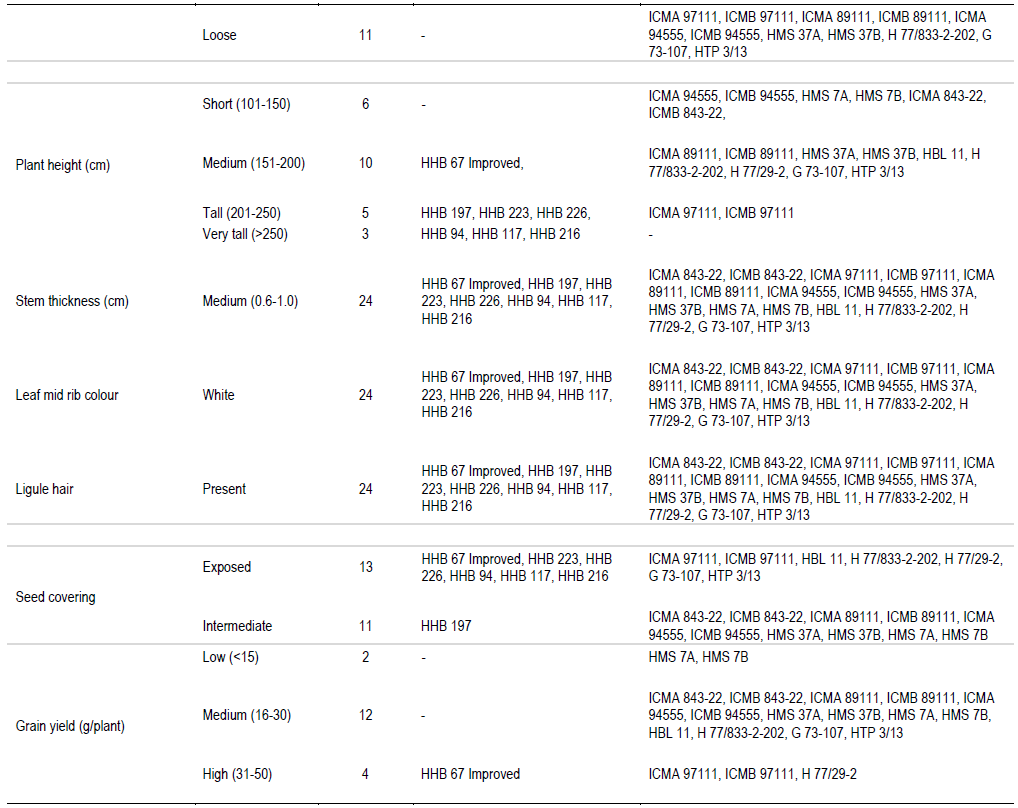
In the present study, sheath pubescence was absent in all the genotype except HMS 7A and HMS 7B. On the basis of number of productive tillers, genotypes were classified in three categories: low, medium and high, having four, sixteen and four genotypes, respectively. On the basis of anther colour, genotypes were classified in two groups: yellow and brown, all the genotypes had yellow anther colour except HHB 94, ICMB 97111 and G 73-107 which possessed brown anther colour. One hybrid (HHB 67 Improved) and eight parental lines showed absence of nodal pubescence whereas all other fifteen genotypes showed presence of nodal pubescence. Among 24 genotypes of pearl millet under study, eleven expressed green nodal pigmentation, four expressed brown nodal pigmentation, six had red nodal pigmentation, two had purple nodal pigmentation and G 73-107 exhibited whitish nodal pigmentation. All the genotypes showed green internode pigmentation except ICMA 97111 and ICMB 97111 which exhibited purple internode pigmentation. All the genotypes showed complete ear head exertion type except HMS 37A and HMS 37B (partial ear head exertion). Only HHB 117 had brown spikelet glum colour whereas all the other genotypes of pearl millet under study were having green spikelet glum colour. HHB 67 improved and its male parent H77/833-2-202 showed purple bristle colour whereas HHB 197, HHB 223, HHB 226, HHB 216 and their male parent HBL 11 and HTP 3/13 had brown bristles. Spike shape classified all the genotypes into four categories: conical, cylindrical, candle and lanceolate. H 77/833-2-202 had lanceolate spike shape, five genotype showed candle spike shape, another five showed cylindrical spike shape and the remaining thirteen were conical in spike shape. Spike tip sterility was present in sixteen genotypes and absent in the remaining eight genotypes. Nine genotypes showed compact spike density, four showed semi compact and the remaining eleven genotypes had loose spike.
Information outlined in Table 2 can effectively be used to find out distinct features of pearl millet genotypes. For example, HHB 117 was the only genotype having brown spikelet glum colour. The HMS 7A and HMS 7B were the only genotype having sheath pubescence. The HHB 94, ICMB 97111 and G 73-107 had brown anther colour. Whitish nodal pigmentation is the unique feature of G 73-107 and purple nodal pigmentation is the unique feature of ICMA 97111 and ICMB 97111. It is also found that partial ear head exertion type is only found in HMS 37A and HMS 37B. H 77/833-2-202 was the only genotype having lanceolate spike shape. These are the qualitative characters and they can help in clear cut identification of particular genotype (Gupta et al., 2011).
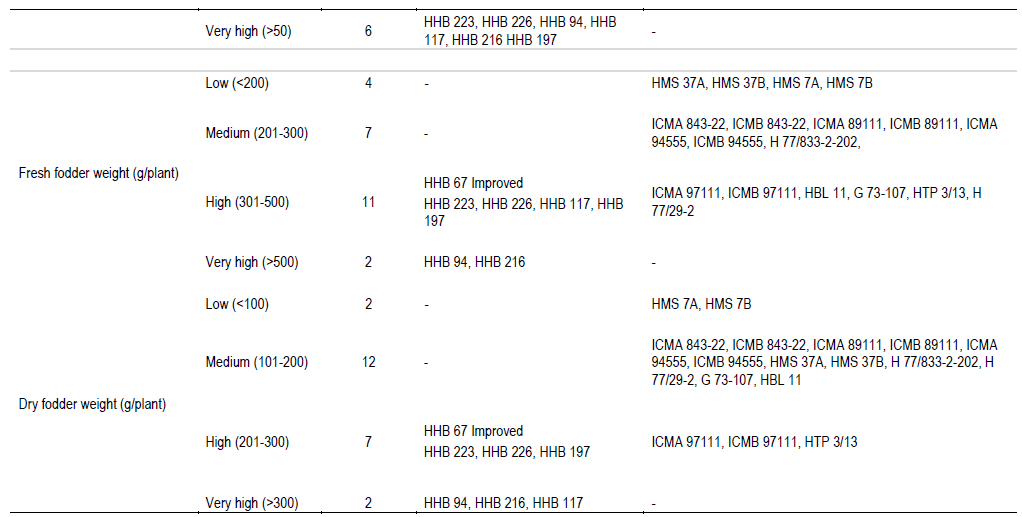
Observations on plant height (short, medium, tall, very tall), leaf length (medium, long, very long), leaf width (narrow, medium, broad), number of nodes (low, medium), spike length (small, intermediate), bristle length (short, medium), grain yield/plant (low, medium, high, very high), fresh fodder weight/plant and dry fodder weight/plant (low, medium, high, very high) were recorded and the data was pooled over the locations. These could be grouped into distinct classes and could be useful for varietal identification and genetic purity testing.
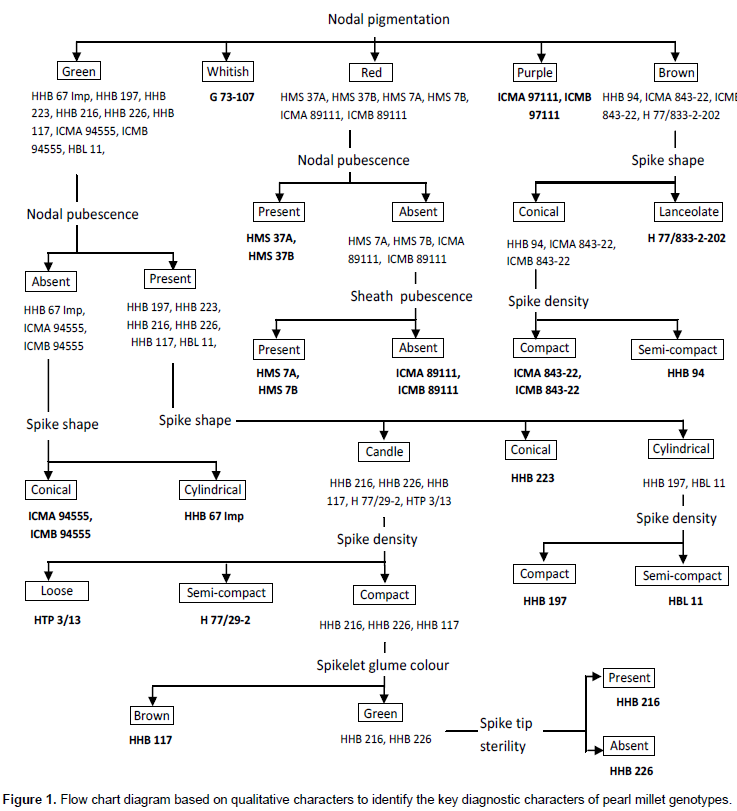
In the Figure 1, flow chart was prepared using qualitative characters to find out distinctive features of all the genotypes (Rana, 2004). Nodal pigmentation was considered as primary character keeping in view the early identification of genotype during plant growth. Nodal pigmentation classified all the genotypes into five groups green, whitish, red, purple and brown. The restorer line G 73-107 could easily be distinguished at early stage of crop growth by having whitish nodal pigmentation. The ICMA 97111 and ICMB 97111 could easily be distinguished by purple nodal pigmentation. Other three nodal pigmentation classes were further separated using nodal pubescence and spike shape. At this level H 77/833-2-202 could easily be distinguished by combination of two characters, that is, nodal pigmentation (brown) and spike shape (lanceolate). In the same way, distinguishing features of all the 24 genotypes were identified through this flow chart based on seven qualitative characters (nodal pigmentation, nodal pubescence, spike shape, spike density, spikelet glume colour, spike tip sterility and sheath pubescence). For example, HHB 94 having a path of three characters spike shape (conical), spike density (semi compact) and nodal pigmentation (brown) is unique for this genotype. It means no other genotype can follow or repeat this path, so this unique path revealed the key diagnostic features of HHB 94. In this flow chart, the CMS line and maintainer line have similar pathway because these two lines are isogenic lines. For example, HMS 7A and HMS 7B have the same key diagnostic features and common path in flow charts.
It may also be concluded that majority of the qualitative/morphological characters found to be dominant in the hybrids were contributed by the male parent. Bristles (present/absent), nodal pubescence, seed covering were found as dominant characters. The study revealed that the morphological characters studied (Table 2), can be used to characterize and catalogued pearl millet genotypes. The key diagnostic characters assigned to the genotypes may also be used for maintaining their seed lots and verification of identity and assessment of varietal purity for seed production, certification and quality control.
The authors have not declared any conflict of interests.