ABSTRACT
The molecular characterization of raw septic tank sewage in the region under study was evaluated using 16S rRNA denaturing gradient gel electrophoresis (DGGE). Raw effluent samples from three septic tanks in the Delta and Edo States region of Nigeria was collected between November 2018 and January 2019 for testing. A composite sample was formed from the three samples collected. The raw sewage sample was sequenced for genomic DNA using Norgen DNA extraction kit to determine the microorganisms present in raw sewage sample. Gene sequence analysis revealed the presence of Methanococcus methanococcus 288171278, Deferribacteres bacterium 291088137, Flavobacteria bacterium 308271278, Bacteroides dorei 671713918, Clostridium difficile 115249003, Kuenenia stuttgartiensis 91203347, Methanosarcina bankeri 827396966, Methanococcus maripaludis 4505076 and Methanobacterium formicicum 693274837, and Desulfitobacterium dichloroeliminans 430782295. The phylogram of the different isolates shows that methane producing bacteria were 7 out of the 13 bacteria isolated; representing 53.8% of the total species occurrence in the sample.
Key words: Gene sequence, bacteria, raw sewage, septic tank.
Sewage, which is liquid waste, is the waste water of a community. It is a combination of water borne wastes from homes, business, health institutions and industries. It may also contain groundwater, surface water and storm water (APHA, 1992). Waste water emanating from sundry human activities may carry pathogenic organisms that can transmit diseases to humans and other animals and contain organic matter that can cause odour and nuisance problems, hold nutrients that may cause eutrophication of receiving water bodies and can lead to ecotoxicology (Doelle, 2001). For reasons of public health and of conservation, man has been forced to develop methods of waste-water storage and treatment which result in the mineralization of the organic components of waste water prior to its discharge into the natural environment. This is usually achieved by an adequate public or community sewerage system. However, such a system is not quite feasible in developing and under developed nations of the world (Rojer, 2002). Hence, individual household sewage disposal system often referred to as septic tank system is the most commonly used domestic waste water disposal method currently being used (Whithers et al., 2014; Schaider et al., 2017; Connelly et al., 2019).
A septic system is an enclosed receptacle designed to collect waste water, segregate settle-able and floatable solids, accumulate and digest organic matter and discharge partially treated effluent. The most widely accepted type is the gravity fed (made of sandcrete block) used by over 40% of the people in Nigeria (Fidelia, 2004). This consists of a septic tank where approximately 54% of the ultimate sewage treatment is accomplished (Robert and Terry, 2004; Schaider et. al., 2017). The septic tank act largely as a settling tank, within which the organic components of the waste water undergo limited aerobic digestion. Effluent from septic tank is as dangerous as raw sewage as it contains effluent concentrations higher than both locally and internationally acceptable limits (Burubai, 2005).
Consequently, attention is being focused on the fermentation of human waste for the production of methane (Zinder, 1993). Methanogenesis is a naturally occurring process that takes place in rice fields and animal digestive tracts, as well as one that can be induced under the right conditions in an artificially constructed anaerobic environment (Lowe et al., 1993; Richards et al., 1994). Both situations utilize the power of methanogens, a bacterial type that can transform organic material into methane through cyclical pathway of production in which intermediate products like acetic acid are formed. Acetate is the ultimate end product of many fermentative pathways and the source of most methane from the anaerobic food chain (Zinder, 1984).
A better understanding of the microbial diversity in septic tank system can improve the stability of the anaerobic process (Bitton, 1999; Connelly et al., 2019). Yet understanding the workings of the process is hampered by the paucity of research papers in the literature (Marti et al., 2013; Ye and Zhang, 2013; Cai et al., 2014; Logares et al., 2015; Newton et al., 2015; Connelly et al., 2019; Numberger et al., 2019). The microbes responsible for the metabolic reactions in the system are the crucial factor in the anaerobic process (Bitton, 1999; Schaider et al., 2017). Understanding and being able to improve our knowledge of the biology of the process as well as the identity of the organisms which promote or inhibit the efficiency of the system is essential to effectively control the start-up and operation of septic tank systems (Schink, 1997; Connelly et al., 2019). Molecular methods like, the PCR-based DGGE technique and sequence analysis have been successfully used to monitor and identify microorganisms within the system (Boon et al., 2002; Odeyemi et al., 2018; Abada et al., 2019; Numberger et al., 2019). The PCR-based DGGE marker construct can effectively be used to study the inherent changes in the microbial population present in the system (Muyzer et al., 1993).
In this work, the diversity, dynamics and degradation potentials of microbial community of the septic tank system was studied using the PCR-based DGGE molecular characterisation technique with the view to further understand and engineer the system in order to optimize its function and utilization.
Study area
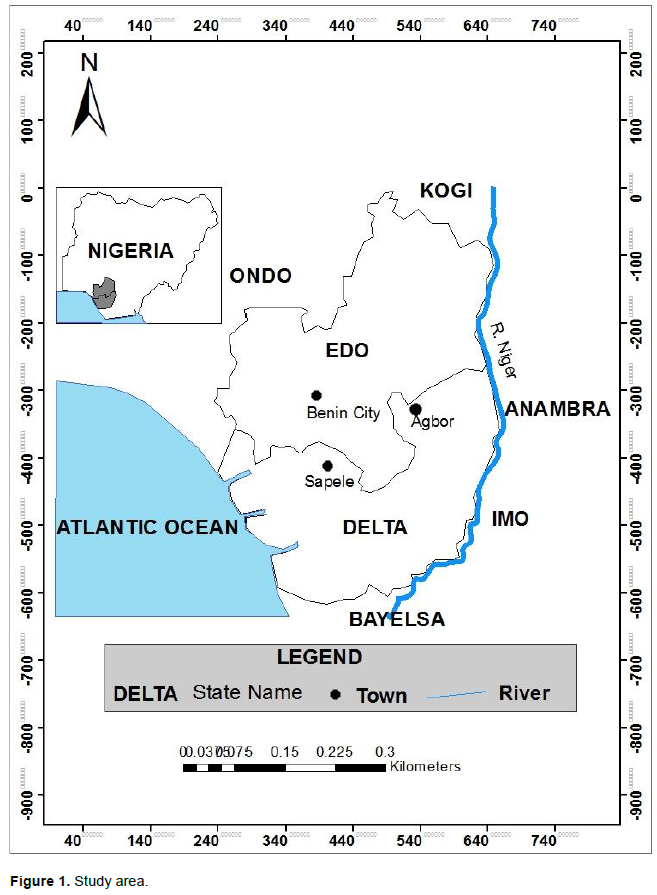
It is a descriptive study carried out in Edo and Delta State, Nigeria in the months of November 2018 and January 2019 with samples analysed at BioSolutions Technologies Laboratory, Akure, Ondo State, Nigeria. The sampling was done in three different locations within Edo and Delta regions of Nigeria (Figure 1); Agbor (A) located 6о 15´ 93´´N and 6о 11´ 59´´E with a population of 222,400 and a land mass of 436 km2, Benin (B) located 6о 20´N and 5о 38´E with a population of 1,471,188 and land mass of 1204 km2 and Sapele (C) located 5о 54´ N and 5о 40´E with a population of 232,000 and a land mass of 394 km2 (NPC, 2006). The major occupation of the people is farming and trading.
Molecular analysis
Methodology was based on PCR and Sanger sequencing analysis (Sanger and Coulson, 1975).
DNA extraction
Ten millilitre of the sewage sample was filtered through 0.22 µm filter pore and then was thawed in a minus 86°C freezer in three consecutive times to release DNA. Further DNA purification was done with Norgen DNA extraction kit. Analysis was done at BioSolutions Technologies Laboratory Akure.
Polymerase chain reaction procedures
Sample was gently vortexed and all solutions after thawing were briefly centrifuged using Eppendorf centrifuge model 5402- Germany. A reaction master mix was prepared by adding the following components (except template DNA) for each 25 µL reaction to a tube at room temperature. The master mix was thoroughly mixed and appropriate volumes were dispensed into PCR tubes or plates. Template DNA (≤500ng/reaction) was added to the individual PCR tubes or wells containing the master mix. Sequence for primers used were PfastBact (Forward): 5' (GGA TTA TTC ATA CCG TCC CA) 3’ PfastBact (Reverse): 5' (CAA ATG TGG TAT GGC TGA TT) 3’ EUB (Forward): 5' (GCA CAA GCG GTG GAG CAT GTGG); 3’EUB (Reverse): '5' (GCC CGG GAA CGT ATT CAC CG) 3’. The PCR cycle started with an initial denaturation step at 94°C for 5 min. This was followed by 30 cycles of denaturation at 94°C for 30 s, annealing at 55°C for 30 s, extension at 72°C for 30 s, and a final extension at 72°C for 5 min that was followed by cooling to 4°C. Few microliters of the samples were run on a 1% agarose gel at 90 V for 30 min in order to verify amplification. The entire PCR reaction was loaded onto a 1% agarose gel and the correct band size (approximately 1500 bp) was excised. Data acquisition was performed during the annealing/extension step.
Acrylamide gel procedure
After determining the percentage gel to be poured, the separation gel layer was used after adding the necessary volumes of reagents to a 100 ml flask (gel will begin polymerizing after adding per sulfate and TEMED; these were added just before pouring the gel). Solution was mixed well by gently swirling the flask. The solution was then quickly poured (or pipetted) into the gel casting apparatus and allowed to polymerize. A flat interface between the separating and stacking layers was ensured. The separation layer was gently overlaid with water or n-butanol after pouring the separation layer. The stacking layer was poured on top of the solution gel, after the separating gel layer has been polymerized; the water or butanol overlay was then decanted. The stacking layer was prepared as prescribed. Necessary volumes of reagents were sequentially added to a flask. Solution was mixed well by gently swirling the flask then quickly pouring (or pipetting) the solution on top of the separation layer. Comb was inserted and layer allowed to polymerize completely before removing comb. Gel was then placed in the electrophoresis apparatus at 75 V.
Preparation of samples
Sixty microlitre of 30, 60, 90, and 100% urea and NaOH (denaturants) was dispensed into Eppendoff tube then 4 microlitre of amplicons was pipetted into them and homogenized thoroughly. Samples were incubated at room temperature for 30 min. PCR was then redone. Pipetted amplicons were put into acrylamide gel and the electrophoresis procedures ran for at least an hour.
Running the gel
The 10x buffer concentrate was diluted (1:9) to make a 1X solution of the buffer. To make 1 L of 1X electrophoresis buffer, 100 mL of the buffer concentrate was added to 900 mL of distilled water. The appropriate volume was then added to the electrophoresis apparatus and ran according to the manufacturer's instructions.
DNA sequencing
Sequence analysis from resultant nucleotides base pairs was performed using BLAST analysis by direct blasting on American data base http://blast.ncbi.nlm.nih.gov. For every set of isolate, a read was Basic Local Alignment Search Tool (BLAST) and the resultant top hits with minimum E-score for every BLAST result showing species name was used to name the specific organism. Sequencing result in FASTA format and corresponding ID after BLAST analysis on NCBI website. BLAST is a computer algorithm program used for comparing nucleotide or amino acid sequences of DNA and/or RNA with available sequences of online database of the National Center of Biotechnology Information (NCBI) and calculates statistical significance (Donkor et al., 2014; NCBI, 2017, 2020).
The result of the molecular characterisation of the raw sewage sample obtained from septic tank using 16S rRNA DGGE (Figure 2) show distinct bands on sodium dodecyl sulphate (SDS) ranging from 18.5 to 100 kDa. Two primers were used representing the two lanes of bands. Each band on the lanes represents a gene from a bacterium.
Result from the PCR analysis of raw domestic sewage sample obtained from the various locations sampled and the subsequent blasting of the sequences using NCBI Blast Online shows the presence of mainly anaerobic and facultative anaerobic methanogens. Methanococcus methanococcus, Deferribacteres bacterium (partial genome 291088137), Flavobacteria bacteria , Bacteroides dorei (CP 008741) Clostridium difficile (AM180355), Kuenenia stuttgartiensis, (CT573073), Zymonas mobilis (AE008692), Methanosarcina bankeri (CP00746), Methanococcus maripaludis (BX950229), Bacteroides metaiotamicron (CAE015928), Methanobacterium formicicom (CP006933), Desulfitobacterium dichloroeliminans (CP003344) and Desulfobacterium sp. (FR695868) are the different species identified from the study. The phylogram of the different isolates (Figure 3) shows that methane producing bacteria were 7 out of the 13 bacteria isolated representing 53.8% of the total species occurrence in the sample.
The result from the PCR analysis of raw domestic sewage sampled in this study and the subsequent blasting of the sequences using NCBI Blast Online shows the presence of mainly anaerobic and facultative anaerobic organisms especially of the methanogen group. Methanococcus methanococcus, Deferribacteres bacterium (partial genome 291088137), Flavobacteria bacteria, Bacteroides dorei (CP 008741), Clostridium difficile (AM180355), Kuenenia stuttgartiensis (CT573073), Zymonas mobilis (AE008692), Methanosarcina bankeri (CP00746), Methanococcus maripaludis (BX950229), Bacteroides metaiotamicron (CAE015928), Methanobacterium formicicum (CP006933) Desulfitobacterium dichloroeliminans (CP003344) and Desulfobacterium sp. (FR695868) are the different species identified from the analysis. Zabranska and Pokorna (2018), as well as Connelly et al. (2019) reported a microbial community underpinned by anaerobic degrading methanogens. The phylogram of the different isolates shows that methane producing bacteria were 7 out of the 13 bacteria isolated (53.8%) of the total species occurrence in the sample. This provided an overall picture of the microbial community present the septic tank. This result is in consonance with the work of Connelly et al. (2019) who stated that the septic system appeared to be underpinned by microbial communities that had the potential to support complete degradation of organics by methanogenic anaerobic digestion. Sequence analysis of the 16S rRNA gene has been widely used to identify bacterial species and perform taxonomic studies (Petti, 2007; Odeyemi et al., 2018; Abada et al., 2019; Numberger et al., 2019). Unfortunately, 16S rRNA hyper variable regions exhibit different degrees of sequence diversity, and no single hyper variable region is able to distinguish among all bacteria. Bacterial activity can be divided into two major distinct phases in the anaerobic digestion; namely acidogenesis (during which acid forming bacteria reduce complex organic matter to organic acids) and methanogenesis (during which specific methanogens may convert the acetate into methane and carbon (IV) oxide). As methane production is an end product, it can also be used to predict septic tank efficiency. Thus, it is important to be able to define the methanogenic species present in these reactors (Smith et al., 1989). Similar organisms were obtained in a study carried out by Garrity and Holt (2001) where they found methanogens in anaerobic sediments and anaerobic sewage sludge bioreactors. Species within this family use acetate as their sole energy source, which is metabolised into methane and carbon IV oxide. The diversity of the methanogenic population depends mainly on the composition of the substrate (Levesque and Guiot, 2004), changes in temperature, pH stability (Odeyemi et al., 2018) and indicators as well as the solids retention time (Casserly and Erijman, 2003; Schaider et al., 2017). Clay soils are known to have higher retention time with a hydraulic conductivity ratio of 1 × 10-11 to 4.7 × 10-9 mm.
PCR analysis also confirms the presence of Desulfitobacterium dichloroeliminans (CP003344) and Desulfobacterium sp. (FR695868) which are sulphate-reducing bacteria in this study. This aligned to the hybridization analysis of 16S rDNA DGGE to investigate the diurnal behaviour of sulphate reducing bacteria in biofilms from an activated sludge basin of a wastewater treatment plant by Teske et al. (1998) were profiles showed that Desulfobulbus and Desulfovibrio populations came up at the onset of the sulphate reduction.
The use of traditional microbiological techniques in determining population structures and characteristics is limited as it has been shown that many organisms are not readily cultured on selective media (Briones and Raskin, 2003). A better understanding of the diversity of the methanogenic bacteria in bioreactors can improve the anaerobic process stability (Bitton, 1999). The methanogens are responsible for the terminal metabolic reactions in a bioreactor and are considered to be the key players in the anaerobic process. The ability to monitor methanogens and understand their ecology is essential to effectively control the start-up and operation of anaerobic digesters in general and the septic tank specifically (Schink, 1997). Finally, certain phylogenetic groups like the methanogens or the methanotrophs exhibit a restricted metabolic potential, which is determined by characteristic functional genes. The detection of these specific genes for example methanol dehydrogenase structural genes of methanotrophs has been used to verify results obtained by the 16S rRNA sequence analysis (Holmes et al., 1995).
The use of septic tank sewage system is an age long practice that is gaining more and more relevance in recent years due to its ease of use, affordability and efficiency. For the system to perform optimally, its operation and utilization techniques must be well studied. Microorganisms particularly bacteria play an important role in the digestion of septic tank constituents into less toxic compounds which the soil can receive and further act upon. The use of traditional microbiological techniques in determining population structures and characteristics is limited as it has been shown that many organisms are not readily cultured on selective media, hence, the need to utilize molecular techniques in identifying and characterizing microbial species.
The phylogram of the different isolates from this study shows that methane producing bacteria were 7 out of the 13 bacteria isolated (53.8%) of the total species occurrence in the sample, while sulphate-reducing and ammonium-oxidizing bacteria accounted for the remaining 6 isolates representing 46.2% of the total bacteria population. This provided an overall picture of the microbial community present the septic tank. A better understanding of the diversity of the methanogenic bacteria in bioreactors can improve the anaerobic process stability; since methanogens are the most occurring species in many anaerobic digesters.
The methanogens are responsible for the terminal metabolic reactions in a bioreactor and are considered to be the key players in the anaerobic process. The ability to monitor methanogens and understand their ecology is essential to effectively control the start-up and operation of anaerobic digesters in general and the septic tank specifically. Lastly, the introduction of noxious chemicals in the form of disinfectant and cleaning agents by humans, may have significantly affected the microbial community and dynamics of the septic tank, this is considered to be a crucial factor in the low biodegradation efficiency of the septic system.
The author wish to thank Dr. Emmanuel Imariagbe of the Department of Environmental Toxicology University of Benin, Benin City Edo State who made meaningful contributions to this work and Prof. Lawrence Okoror of Biosolutions Technologies Akure, Ondo State Nigeria for running the molecular aspect of this work.
REFERENCES
|
Abada E, Al-Fifi Z, Al-rajab AJ, Mahdi M, Sharma M (2019). Molecular identification of biological contaminants in different drinking water resources of Jazan region, Saudi Arabia. (2019). Journal of Water and Health 17(4):622-632.
Crossref
|
|
|
|
American Public Health Association (APHA) (1992). Standard methods for the examination of water and waste water. American Public Health Association 18th edition Washington D.C.
|
|
|
|
|
Bitton G (1999). Waste water Microbiology, Second Edition, Wineglass, NY. p. 139.
|
|
|
|
|
Boon N, De Windt W, Verstracte W, Top EM (2002). Evaluation of nested PCR-DDE with group specific 16S rRNA primers for the analysis of bacterial communities from different waste water treatment plants microbes. Ecology 39:101-112.
Crossref
|
|
|
|
|
Briones A, Raskin L (2003). Diversity and dynamic of microbial communities in engineered environments and their implications for process stability Current Opinion in Biotechnology14:270-276.
Crossref
|
|
|
|
|
Burubai W (2005). Source water pollution abatement and best management practices. In Proceedings of 7th African USA International Conference on Manufacturing Technology pp. 277-282. Port Harcourt, Nigeria (12-14 July).
|
|
|
|
|
Cai L, Ju F, Zhang T (2014). Tacking human sewage microbiome in a municipal wastewater treatment plant. Applied Microbiology and Biotechnology 98:3317-3326.
Crossref
|
|
|
|
|
Casserly C, Erijman L (2003). Molecular monitoring of microbial diversity in an UASB REACTOR International Biodeterioration and Biodegradation 52:7-12.
Crossref
|
|
|
|
|
Connelly S, Pussayanavin T, Randle-Boggis RJ, Wicheansan A, Jampathong S, Keating C, Ijaz UZ, Sloan WT, Koottatep T (2019). Solar septic tank: Next Generation sequencing reveal a effluent microbial community composition as a useful index of system performance. Water 1:2660.
Crossref
|
|
|
|
|
Doelle HW (2001) Biotechnology and Human Development in developing countries www. Ejbiotechnology. Info.
Crossref
|
|
|
|
|
Donkor ES, Dayie NT KD, Adiku TK (2014). Basic local alignment search tool (BLAST) and fast alignment (FASTA). Journal of Bioinformatics and Sequence Analysis 6(1):1-6.
Crossref
|
|
|
|
|
Fidelia N (2004). Source water pollution abatement and best management practices. In Proceedings of 7th Africa-USA international Conference on Manufacturing Technology, pp. 277-282. Port Harcourt, Nigeria (12-14 July).
|
|
|
|
|
Garrity GM, Holt JG (2001). Phylum AII,. Euryarchaeota phy. Nov., In:D.R. Boone, R.W. Castenholz (Eds), Bergey's Manuel of Systemic Bacteriology second ed., Springer, New York, pp. 211-294.
|
|
|
|
|
Holmes AJ, Owens NJP, Murrell JC (1995). Defection of novel marine methanotrophs using phylogenetic and functional gene probes after methane enrichment. Microbiology 141:1947-1955.
Crossref
|
|
|
|
|
Levesque MJ, Guiot SR (2004). Positioning of Methanosaeta and Methanosarcina sp. within the microstructure of carbohydrate-fed anaerobic granules, In: Proceedings of the 10th Anaerobic Digestion Conference, Montreal Canada pp. 1934-1937.
|
|
|
|
|
Logares R, Mangot J-F, Massana R (2015). Rarity in aquatic microbes: Placing protists on the map. Research in Microbiology 166:831-841.
Crossref
|
|
|
|
|
Lowe SE, Jain MK, Zeikus JG (1993). Biology, ecology and biotechnological applications of anaerobic bacteria adapted to environmental stress in temperature, pH, salinity or substrates. Microbiological Reviews 57:451-509.
Crossref
|
|
|
|
|
Marti E, Jofre J, Balcazar JL (2013). Prevalence of antibiotic resistance genes and bacterial community composition in a river influenced by a wastewater treatment plant. PLoS ONE 8:e78906
Crossref
|
|
|
|
|
Muyzer G, Waal EC de, Uitterlinden AG (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA Applied and Environmental Microbiology 59:695-700.
Crossref
|
|
|
|
|
National Centre for Biotechnology Information (NCBI) (2017).Pubchem Compound Database (ID) Sept 16, 2017.
|
|
|
|
|
National Centre for Biotechnology Information (NCBI) (2020). Pubchem Compound Database (ID) May 27, 2020.
|
|
|
|
|
Newton RJ, McLellan SL, Dila DK, Vineis JH, Morrison HG, Murat Eren A (2015). Sewage reflects the microbiomes of human population. mBio 6(2):e02574-14.
Crossref
|
|
|
|
|
Numberger D, Ganzert L, Zoccarato L, Muhldorfer K, Sauer S, Grossart H-P, Greenwood AD (2019). Characterization of bacterial communities in wastewater with enhanced taxonomic resolution by full-length 16S rRNA sequencing. Nature 9:9673.
Crossref
|
|
|
|
|
Odeyemi AT, Ayantola KJ, Peter S (2018). Molecular characterization of bacterial isolates and physicochemical assessment of well water samples from hostels at Osekita, Iworoko-Ekiti, Ekiti State. American Journal of Microbiology Research 6(1):22-32.
Crossref
|
|
|
|
|
Petti CA (2007). Detection and Identification of microorganisms by gene amplification and sequencing. Clinical Infectious Diseases 44:1108-1114.
Crossref
|
|
|
|
|
Richards BK, Herndon HG, Jewell WJ, Cummings RJ, White TE (1994). In situ methane enrichment in methanogenic energy crop digesters. Biomass and Bioenergy 6(4):275-282.
Crossref
|
|
|
|
|
Robert W, Terry B (2004). Septic tank operations. USDA-104. GPO Washington DC., p. 76
|
|
|
|
|
Rojer RN (2002). Introduction to Environmental analysis. New York: John Wiley and Sons Ltd. pp 313.
|
|
|
|
|
Schaider LA, Rodgers KM, Rudel RA (2017). Review of organic wastewater compounds concentrations and removal in onsite wastewater treatment systems. Environmental Science Technology 51:7304-7317.
Crossref
|
|
|
|
|
Schink B (1997). Energetics of syntrophic cooperation in methanogenic degradation. Microbiological and Molecular Biology Review 94:262-280.
Crossref
|
|
|
|
|
Smith T, Ince M (1989). Septic System Density and Ground Water Contamination in Illinois: A survey of state and local regulation. NTIS Report PB. 89-178545.
|
|
|
|
|
Sanger F, Coulson AR (1975). A rapid method for determining sequence in DNA by primed synthesis with DNA polymerase. Journal of Molecular Biology 25:(3) 441-448.
Crossref
|
|
|
|
|
Teske A Ramsing NB, Habicht K, Fukui MK, Uver J. Jergensen BB, Cohen Y (1998). Sulphate -reducing bacteria and their activities in cyanobacteria mats of solar lake (Sinai Egypt). Applied and Environmental Microbiology 64:2943-2951.
Crossref
|
|
|
|
|
Whithers PJA, Jordan P, May L, Jarvie HP, Deal NE (2014). Do septic tank systems pose a hidden threat to water quality? Frontiers in Ecology and the Environment 12:123-130.
Crossref
|
|
|
|
|
Ye L, Zhang T (2013). Bacterial communities in different sections of a wasterwater treatment plant revealed by 16S rDNA 454 pyrosequencing. Applied Microbiology and Biotechnology 97:2681-2690.
Crossref
|
|
|
|
|
Zabranska J, Pokorna D (2018). Bioconversion of carbon dioxide to methane using hydrogen and hydrogenotrophic methanogens. Biotechnology Advances 36:707-720.
Crossref
|
|
|
|
|
Zinder SH (1984). Microbiology of anaerobic conversion of organic wastes to methane: Recent developments. News 50:294-298.
|
|
|
|
|
Zinder SH (1993). Physiological ecology of methanogens. In Ferry, J.G (ed) Methanogenesis: Ecology, Physiology, Biochemistry and Genetics. New York: Chapman and Hall. pp. 128-206.
Crossref
|
|